Using the tabular output generated by uc_process, this function will build a graph to
visualize the results. Each function configuration will output a bespoke ggplot. Theming can
be adjusted by the user after the graph has been output using + theme(). Most graphs can
also be made interactive using make_interactive_squba()
Usage
uc_output(
process_output,
output_col,
filter_variable = NULL,
large_n = FALSE,
large_n_sites = NULL
)Arguments
- process_output
tabular input || required
The tabular output produced by
uc_process- output_col
string || defaults to
NULLThe name of the numerical variable from
process_outputthat should be used to generate the plot. This input is required for all checks EXCEPTMulti Site, Anomaly Detection, Longitudinal- filter_variable
string or vector || defaults to
NULLA string or vector with variable names that should be the focus of the analysis. Only a single value is accepted for the following checks:
Multi Site, Anomaly Detection, LongitudinalSingle Site, Anomaly Detection, Longitudinal
Multiple values are accepted for:
Multi Site, Exploratory, Cross-SectionalMulti Site, Exploratory, Longitudinal
- large_n
boolean || defaults to
FALSEFor Multi-Site analyses, a boolean indicating whether the large N visualization, intended for a high volume of sites, should be used. This visualization will produce high level summaries across all sites, with an option to add specific site comparators via the
large_n_sitesparameter.- large_n_sites
vector || defaults to
NULLWhen
large_n = TRUE, a vector of site names that can add site-level information to the plot for comparison across the high level summary information.
Value
This function will produce a graph to visualize the results
from uc_process based on the parameters provided. The default
output is typically a static ggplot or gt object, but interactive
elements can be activated by passing the plot through make_interactive_squba.
For a more detailed description of output specific to each check type,
see the PEDSpace metadata repository
Examples
#' Source setup file
source(system.file('setup.R', package = 'unmappedconcepts'))
#' Create in-memory RSQLite database using data in extdata directory
conn <- mk_testdb_omop()
#' Establish connection to database and generate internal configurations
initialize_dq_session(session_name = 'uc_process_test',
working_directory = my_directory,
db_conn = conn,
is_json = FALSE,
file_subdirectory = my_file_folder,
cdm_schema = NA)
#> Connected to: :memory:@NA
#' Build mock study cohort
cohort <- cdm_tbl('person') %>% dplyr::distinct(person_id) %>%
dplyr::mutate(start_date = as.Date(-15000), # RSQLite does not store date objects,
# hence the numerics
end_date = as.Date(20000),
site = ifelse(person_id %in% c(1:6), 'synth1', 'synth2'))
#' Execute `uc_process` function
#' This example will use the single site, exploratory, cross sectional
#' configuration
uc_process_example <- uc_process(cohort = cohort,
multi_or_single_site = 'single',
anomaly_or_exploratory = 'exploratory',
time = FALSE,
omop_or_pcornet = 'omop',
uc_input_file = uc_input_file_omop) %>%
suppressMessages()
#> ┌ Output Function Details ─────────────────────────────────────┐
#> │ You can optionally use this dataframe in the accompanying │
#> │ `uc_output` function. Here are the parameters you will need: │
#> │ │
#> │ Always Required: process_output │
#> │ Required for Check: output_col │
#> │ │
#> │ See ?uc_output for more details. │
#> └──────────────────────────────────────────────────────────────┘
uc_process_example
#> # A tibble: 2 × 13
#> site variable total_rows total_pt unmapped_rows unmapped_pt median_all_with0s
#> <chr> <chr> <int> <int> <int> <int> <dbl>
#> 1 comb… visit t… 1627 12 0 0 0
#> 2 comb… hyperte… 58 12 13 1 0
#> # ℹ 6 more variables: median_all_without0s <int>, median_site_with0s <dbl>,
#> # median_site_without0s <int>, unmapped_row_prop <dbl>,
#> # unmapped_pt_prop <dbl>, output_function <chr>
#' Execute `uc_output` function
uc_output_example <- uc_output(process_output = uc_process_example,
output_col = 'unmapped_row_prop')
uc_output_example
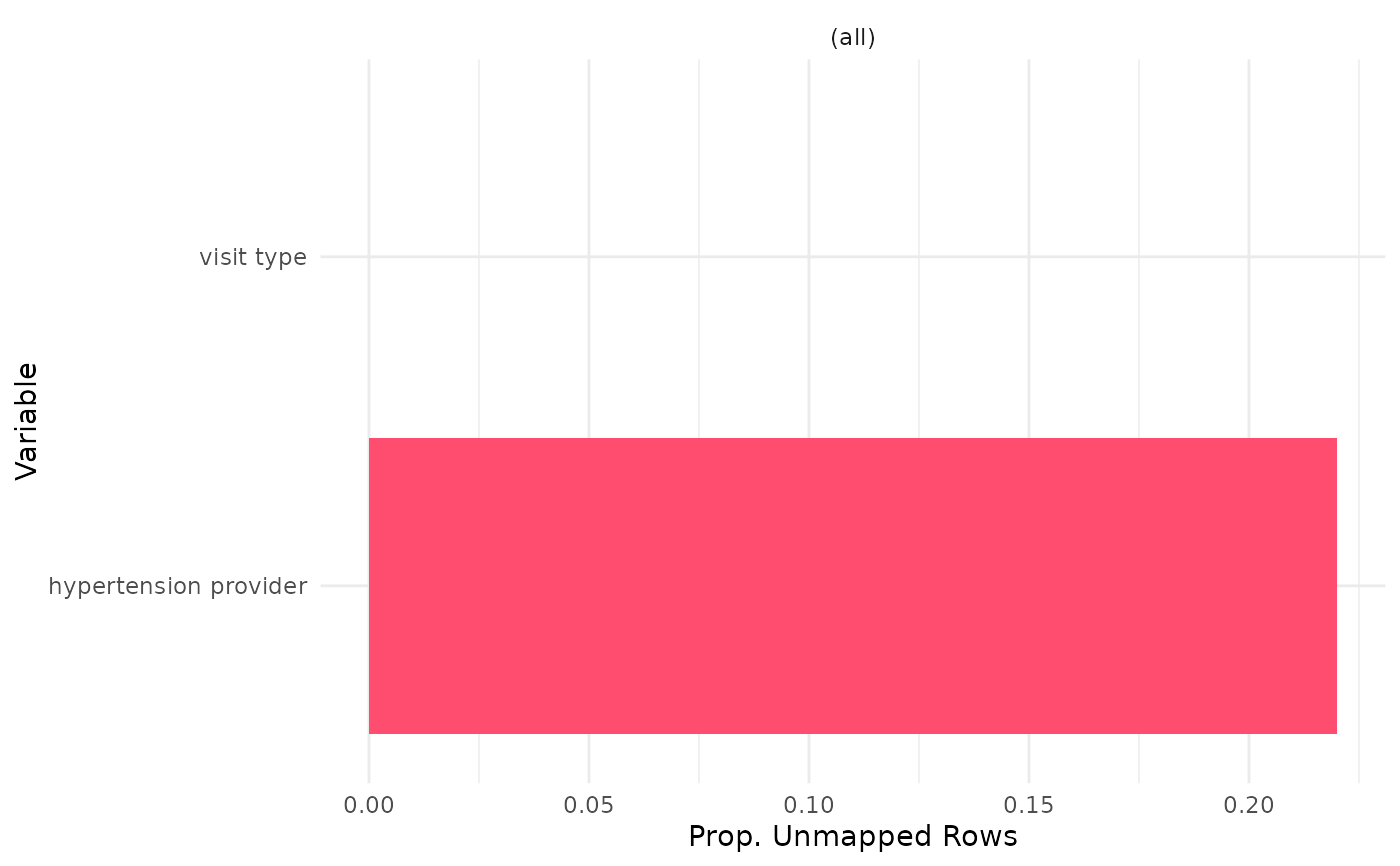 #' Easily convert the graph into an interactive ggiraph or plotly object with
#' `make_interactive_squba()`
make_interactive_squba(uc_output_example)
#' Easily convert the graph into an interactive ggiraph or plotly object with
#' `make_interactive_squba()`
make_interactive_squba(uc_output_example)